Introduction

Phylogenetic trees are a way to visualize evolutionary relationships between species for a factor or gene of particular interest [1]. The branching of the trees demonstrate how related the organisms are to each other. The branch point indicates a most common recent ancestor between species. Species that are placed together closer after a branch point indicate a closer relationship.
MegaX is a free software program to generate phylogenetic trees [2]. FASTA sequences containing the string of amino acids that make up the protein can be inputted to the software for each model organism. The program will then align the sequences to construct a phylogenetic tree.
results
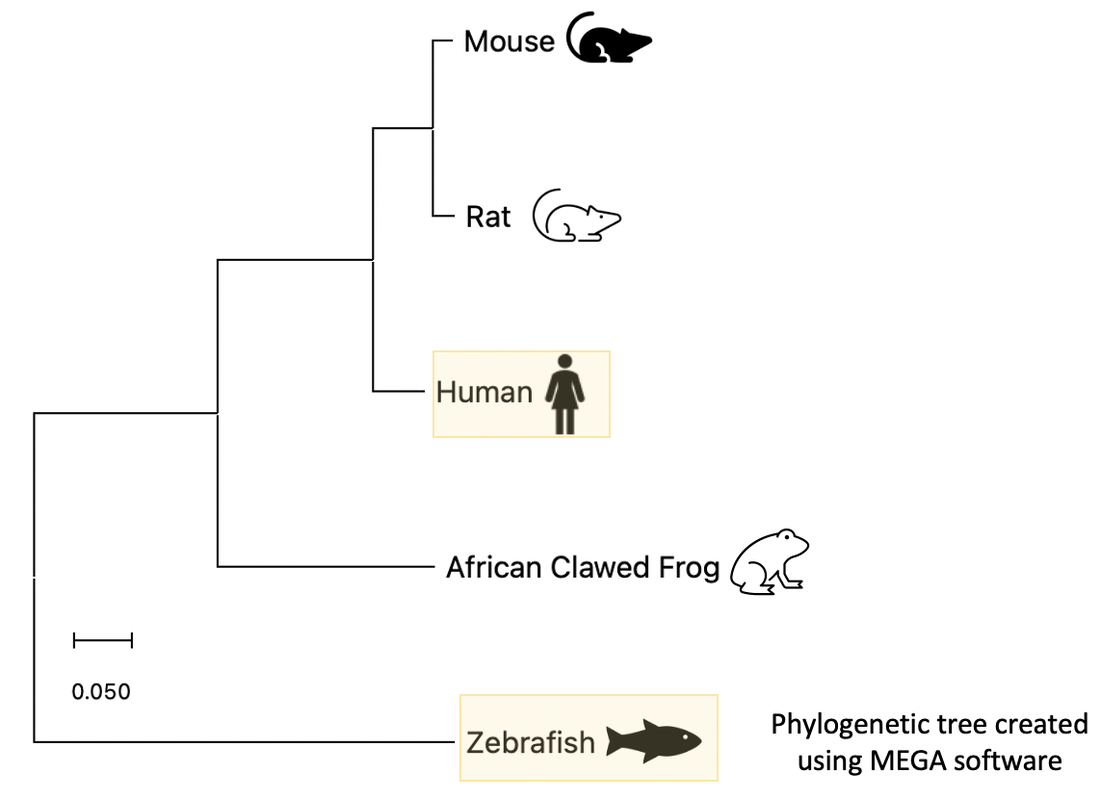
Phylogenetic tree for the COL3A1 protein
Conclusion
This tree demonstrates phylogeny for the COL3A1 protein and was assembled using maximum likelihood in MegaX. There is the closest relationship of COL3A1 sequences seen in humans, mice, rats, african clawed frog, and zebrafish. Many of the species listed would be effective model organisms to study COL3A1, and the relationships seen here could be used in the decision making process.
References
[1] Khan Academy. (2016). Phylogenetic trees | Evolutionary tree (article). Khan Academy. https://www.khanacademy.org/science/ap-biology/natural-selection/phylogeny/a/phylogenetic-trees
[2] https://www.megasoftware.net/
[2] https://www.megasoftware.net/
This web page was produced as an assignment for Genetics 564, a capstone course at UW-Madison